Kun Zhou1,2,*, James C. Kosmopoulos2,3, Etan Dieppa Colón2,3, Peter John Badciong2, Karthik Anantharaman2,4,5,*
1 State Key Laboratory of Marine Geology, Tongji University, Shanghai, China
2 Department of Bacteriology, University of Wisconsin–Madison, Madison, WI, USA
3 Microbiology Doctoral Training Program, University of Wisconsin-Madison, Madison, WI, USA
4 Department of Integrative Biology, University of Wisconsin–Madison, Madison, WI, USA
5 Department of Data Science and AI, Wadhwani School of Data Science and AI, Indian Institute of Technology Madras, Chennai, India
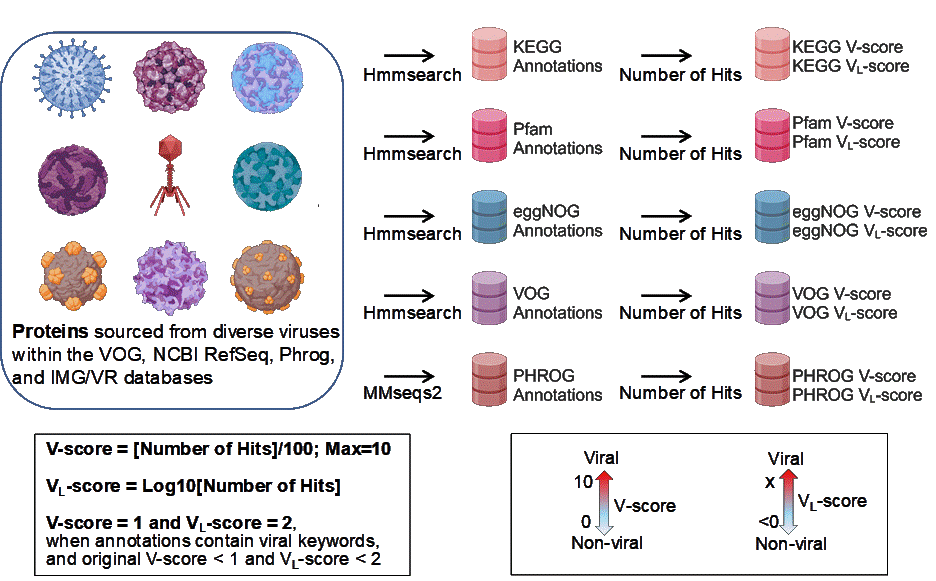
Abstract: Viruses are key drivers of microbial ecology evolution, yet their study is hindered due to challenges in culturing. Traditional gene-centric methods, which focus on a few hallmark genes like for capsids, miss much of the viral genome, leaving key viral proteins and functions undiscovered. Here, we introduce two powerful annotation-free metrics, V-score and VL-score, designed to quantify the “virus-likeness” of protein families and genomes and create an open-access searchable database, ‘V-Score-Search’. By applying V- and VL-scores to public protein databases, we link 19-59% of protein families with viruses representing a 5-8x increase over current estimates. These metrics outperform existing approaches, enabling high efficiency in detection of viral genomes, prophages, and host-derived auxiliary viral genes (AVGs) from fragmented sequences. Remarkably, we identify up to 17 times more AVGs dominated by non-metabolic proteins of unknown function. This innovation unlocks new insights into virus signatures and host interactions, with wide-ranging implications from genomics to biotechnology.
Full article: https://www.nature.com/articles/s41467-026-72028-0